Helicobacter pylori (H. pylori) infection is the most widespread infectious-contagious disease worldwide, reaching a prevalence of 50–80% in developing countries. Chronic infection is considered the main cause of chronic gastritis and has been related to other diseases, such as peptic ulcer, gastric mucosa-associated lymphoid tissue lymphoma, and gastric cancer. The most common treatment is with eradication regimens that utilize three or four drugs, including a proton pump inhibitor (PPI) and the antibiotics, clarithromycin and amoxycillin or metronidazole. Empiric antibiotic use for eradicating the bacterium has led to a growing resistance to those drugs, reducing regimen efficacy and increasing costs for both the patient and the healthcare sector. In such a context, the development of noninvasive next-generation molecular methods holds the promise of revolutionizing the treatment of H. pylori. The genotypic and phenotypic detection of the resistance of the bacterium to antibiotics enables personalized treatment regimens to be provided, reducing costs and implementing an antibiotic stewardship program.
The aims of the present narrative review were to analyze and compare the traditional and next-generation methods for diagnosing H. pylori, explain the different factors associated with eradication failure, and emphasize the impact of the increasing antibiotic resistance on the reversal and prevention of H. pylori-associated diseases.
La infección por Helicobacter pylori (H. pylori) es la enfermedad infecto-contagiosa más diseminada a nivel mundial, alcanzando una prevalencia del 50–80% en países en vías de desarrollo. La infección crónica es considerada como la principal causa de gastritis crónica, y se ha relacionado con otras enfermedades como úlcera péptica, linfoma de tejido linfoide asociado a mucosa gástrica y cáncer gástrico. El tratamiento más común son los esquemas de erradicación que utilizan tres o cuatros fármacos, entre ellos un inhibidor de bomba de protones (IBP) y dos antibióticos, claritromicina y amoxicilina o metronidazol. El uso empírico de antibióticos para la erradicación de la bacteria ha propiciado una creciente resistencia a dichos fármacos, disminuyendo la eficacia de los esquemas y aumentando los costos para el paciente y sector salud. Es por esto que el desarrollo de métodos moleculares no invasivos de siguiente generación promete ser una herramienta que revolucione la terapéutica en H. pylori. La detección genotípica y fenotípica de resistencia a antibióticos de la bacteria permite brindar esquemas personalizados de tratamiento, disminuir costos e implementar un programa de administración de antibióticos.
Los objetivos de esta revisión narrativa son analizar y comparar los métodos diagnósticos tradicionales y de siguiente generación para el diagnóstico de H. pylori, explicar los diversos factores asociados a falla de erradicación y puntualizar sobre el impacto de la creciente resistencia a antibióticos sobre la reversión y prevención de enfermedades asociados a H. pylori.
Helicobacter pylori (H. pylori) is a spiral-shaped, flagellated Gram-negative bacterium that specifically colonizes the human gastric mucosa. H. pylori infection is the most widespread infectious-contagious disease worldwide. In 2015, an estimated 4,400,000 persons, more than 50% of the world population, were infected by the bacterium and prevalence reached 80% in developing countries1. Even though the infection rate varies, in the past three decades, the number of infected persons has increased due to population growth, as well as to eradication treatment failure2. In addition, the reinfection rate after successful treatment is estimated at 2%3.
Therefore, carrying out a narrative review was decided upon, whose aims were to analyze and compare traditional and next-generation methods for diagnosing H. pylori, explain the different factors associated with eradication failure, and emphasize the impact of increasing antibiotic resistance on the reversal and prevention of H. pylori-associated disease. The methodology employed to develop the review was a bibliographic search, independently conducted by two researchers, utilizing the PubMed search engine for clinical trials, systematic reviews, meta-analyses, and clinical guidelines published within the time frame of 1990 and 2021. The search was focused on next-generation molecular methods, antimicrobial resistance, novel treatments, and antibiotic administration regimens for H. pylori, using the terms (Helicobacter pylori [MeSH]) AND (next generation sequencing), (Helicobacter pylori [MeSH]) AND (antibiotic* OR antimicrobial*) AND (resistance), (Helicobacter pylori [MeSH]) AND (treatment) and (Helicobacter pylori [MeSH]) AND (treatment) AND (failure).
Prevalence of H. pylori varies greatly in the different countries and regions of the world. It is higher in Africa (79.1%), Latin America and the Caribbean (63.4%), and Asia (54.7%), whereas the regions of lower prevalence are North America (37.1%) and Oceania (24.4%)1. Although the infection route has not been completely defined, the prevalent hypothesis suggests fecal-oral transmission, in which the infected mother or grandmother vertically transmits the bacterium to the child in the first years of life4. The differences in prevalence can be explained by different degrees of access to clean water, urbanization, sanitation, and socioeconomic status, among other factors2,5.
In the co-evolutionary analysis of the bacterium and its adaptation in the Americas, the genomic sequencing of 723 varieties of H. pylori indicated that the bacterium entered the continent by way of Mexico6. In the Mexican population, in particular, the reported national seroprevalence average of H. pylori is 66%7. Even though prevalence is high, a decrease has been found, especially in children and young adults, which is mainly thought to be due to an improved national socioeconomic condition8.
In 2017, the World Health Organization (WHO) declared that H. pylori was one of the 12 “priority pathogens” because of its increasing resistance to antibiotics, and placed it in the high priority category, given its status as a public health threat, and as a group 1 carcinogen, according to the International Agency for Research on Cancer (IARC)9–11. The majority of infections are asymptomatic, in up to 80% of cases, and there is actually no urgency for eradicating H. pylori12,13. When symptomatic, the infection is associated with peptic ulcer disease in 5–10% of cases, dyspepsia in 1–10%, non-cardia gastric cancer in <1%, and mucosa-associated lymphoid tissue (MALT) lymphoma in <0.1%12,14,15. Despite the low percentage of patients that develop neoplasia, gastric cancer is the third cause of cancer-related death worldwide, responsible for 750,000 deaths annually. Ninety-percent of non-cardia gastric cancers are attributable to H. pylori10,16,17. Extraintestinal association of the infection, such as iron deficiency, vitamin B12 deficiency, and idiopathic thrombocytopenic purpura, among others, is also well described12,18. Interestingly, the infection is a protective factor against the development of gastroesophageal reflux disease, Barrett’s esophagus, and esophageal adenocarcinoma10. Indeed, some authors question the pathogenic role of the bacterium, considering it to be part of the normal human microbiota, acting as a commensal or even a symbiont14.
Current status quo of eradication treatmentThe causes of H. pylori infection eradication failure can be divided into factors related to the host, related to the bacterium, and associated with the healthcare system.
Regarding the bacterium-associated factors, antimicrobial resistance is the most common. Antibiotic resistance rates are estimated at 10–34% for clarithromycin, 11–30% for levofloxacin, and 23–56% for metronidazole17. Geographically, antibiotic resistance rates vary widely, and that diversity has been on the rise for years16. In Mexico, resistance to clarithromycin is reported at 13%, to metronidazole at 60%, to amoxicillin at 4%, to tetracyclines at 2%, and dual resistance to metronidazole/clarithromycin is reported at 13%8. It should be underscored that resistance to amoxicillin and the tetracyclines is extremely rare, even after infection eradication failure with a regimen including those antibiotics, and no resistance to bismuth has yet been described19. Resistant bacteria present with a resistance phenotype, i.e., they can grow in the presence of an antibiotic, and utilize an active mechanism associated with an inheritable mutation to withstand antibiotic-induced stress20.
In addition to the resistance phenotype, two more are associated with failed antibiotic treatment: the tolerance phenotype and the persistence phenotype20. Tolerance is defined as the capacity of a bacterial population to survive exposure to elevated concentrations of an antibiotic, with no modification of the minimum inhibitory concentration (MIC), by slowing down essential bacterial processes. That phenotype can be acquired through exposure to conditions of environmental stress21,22. Persistence is the phenomenon in which a bacterial population exhibits epigenetic traits of inactivity and the absence of metabolic activity, to tolerate antibiotics23,24. Features of the persistence phenotype are the ceasing of cellular activity, the absence of growth or change in concentration in the presence of the antibiotic, and the capacity to rapidly revert to the growth pattern and customary activity once the antibiotic is eliminated and the conditions are optimal for their development20,24,25. The tolerance phenotype and the persistence phenotype are essentially different from the resistance phenotype in that neither of them utilizes an active mechanism to withstand antibiotic-associated stress, tolerant and persistent populations do not grow under drug pressure, and they are not inherited phenotypes20. Different cellular mechanisms are involved to a greater or lesser degree in the phenomena of tolerance and resistance. The main ones are the general response to stress, oxidative tolerance, the bacterial communication system (quorum sensing), the (p)ppGpp signaling system, and the toxin-antitoxin modules26. Explaining the molecular mechanisms behind those processes is beyond the scope and aims of this narrative review.
An important phenotypic characteristic of H. pylori is that, upon activating its stress response mechanisms, it can present with morphologic variability, i.e., it transforms from its active spiral-shaped form to an inactive coccoid-shaped form27,28. The coccoid forms are resistant to gastric acid and antibiotics, they enable immune system evasion, and they are transmitted with greater efficacy. In addition, they have the capacity to remain latent, without losing their viability or virulence factors, enabling chronic and treatment-refractory infection29,30. Added to hampering treatment, the coccoid form is a diagnostic challenge, given that it cannot be cultured, it produces false negatives in 13C-labeled urea breath tests, and can cause confusion in morphologic identification in biopsies31.
To achieve a persistent infection, H. pylori, despite producing a strong immune response, has different mechanisms for evading, interrupting, and manipulating that response, which is both innate and adaptive in the host, consequently preventing the bacterium’s eradication32,33. Some of the mechanisms utilized for evading the innate immune response include the evasion of the recognition and mediation of the Toll-like receptor (TLR) and C-type lectin receptor (CLR) signaling and the use of molecular mimicry34. To prevent recognition by the TLRs, the H. pylori lipopolysaccharides (LPSs) have structural changes that reduce their detection by those receptors, involving acylation (tetra-acylated lipid A) and the absence of phosphate groups at the 1′ and 4′ positions of lipid A35,36. To modulate the TLR-mediated immune response, the bacterium activates TLR9, the intracellular receptor found in dendritic cells, and TLR2, both of which trigger an anti-inflammatory response37. Of the CLRs, one of the best described is the C-type II lectin receptor, known as Dendritic Cell-Specific Intercellular Adhesion Molecule-Grabbing Nonintegrin (DC-SIGN) CD-209, found in macrophages and dendritic cells. The ligands for that H. pylori receptor have fucose residues that activate an anti-inflammatory response32,38. The O antigens of the H. pylori LPSs are modified by fucosyltransferases that produce structures that molecularly mimic Lewis antigens, preventing detection by TLRs by not being recognized as foreign39,40. Evasion of the adaptative immune response is achieved through the VacA, γ-glutamyl transpeptidase (GGT), and CagA virulence factors, which through different molecular mechanisms, together inhibit T lymphocyte proliferation34,41. GGT and VacA are also involved in the differentiation of T cells into regulatory T lymphocytes (Tregs). Said induction is dependent on the age at which the host contracted the infection and is greater when it occurs in infancy, producing tolerance to H. pylori26,42. Another virulence factor associated with tolerance to infection is the outer inflammatory protein A (OipA), a dendritic cell maturation suppression factor, which aids in establishing chronic infection due to tolerogenic programming43.
With respect to the host factors, both genetic and environmental factors have been associated with H. pylori infection eradication failure4. The CYP2C19 polymorphisms are one of the main patient factors involved. Those polymorphisms alter the metabolism, and in turn, the effectiveness of the proton pump inhibitors (PPIs), especially first-generation drugs (omeprazole, lansoprazole)17,44. The CYP2C19 gene has 21 polymorphisms, three of which define the enzyme activity that characterizes the host’s CYP2C19 metabolizer phenotype of PPIs44,45. The rapid metabolizers are the most frequent polymorphism, with a higher prevalence in Whites, African Americans, and Hispanics (57–71 %), and a lower frequency in Asians17. Notably, patients that are homozygous for the CYP2C19*17 allele have been identified as ultra-rapid metabolizers, albeit that variant is not clinically different from the rapid metabolizers45. Even though there is insufficient evidence supporting the genetic study of the CYP2C19 gene polymorphisms in routine clinical practice, to guide H. pylori eradication therapy, those polymorphisms do have clinical implications, given that the rapid and ultra-rapid metabolizers would benefit from a higher dose or more frequent dosing of first-generation PPIs or the use of more recent and stronger PPIs, or if accessible, the use of non-PPI acid suppressants, such as vonoprazan17,44. Other genetic polymorphisms involved are those that affect the intragastric pH, importantly including the IL-1B and MDR1 genes46.
Regarding environmental factors, smoking has most often been associated with H. pylori eradication failure. Smoking increases gastric acid secretion, reduces mucus secretion, and decreases the gastric blood flow, reducing the effectiveness of PPIs and antibiotics4,17,47. Other environmental factors for which there is less evidence of association include age, diabetes, and obesity4.
In addition to the factors presented by the host and by the bacterium, there are healthcare system factors that favor H. pylori infection eradication failure17. A vastly important factor is treatment adherence by the patient; the level of adherence from which there is no further increase in the eradication success rate in H. pylori infections is unknown, but studies state that the minimum of adherence for success varies from 60 to 90% of treatment compliance48,49. Different strategies have been recommended for improving treatment adherence, such as using Smartphone medical applications (apps) or reminders via text messages17,50. The physician should clearly explain the reasoning behind the prescription of the medications, the dosage instructions, and the expected adverse effects, emphasizing the great importance of finishing the complete treatment regimen17. The adverse effects related to H. pylori eradication therapy include nausea, vomiting, diarrhea, and altered sense of taste, whose presence can lead to discontinuing the treatment before its completion and contributing to the lack of adherence and treatment failure51. The costs of the diagnostic approach and treatment can become high, hindering their accessibility for certain sectors of the population19,52.
Conventional methods and novel next-generation molecular methods for diagnosing Helicobacter pylori (Table 1)Rapid urease testThe rapid urease test (RUT) is an indirect test for determining the presence of H. pylori in the gastric mucosa. Unlike serology, it only detects active infection. The test requires obtaining a gastric biopsy sample that is then added to a device, in which the sample binds to urea. The hydrolysis products of urea, ammonia, and carbon dioxide are detected by the presence of the urease enzyme in the bacterium53. The speed of the RUT reaction depends on the bacterial load and temperature. PPI or antibiotic use can cause false negatives. False positives are rare and occur when there are other urease-producing microorganisms present31. Their concentrations must be sufficient to produce a positive result, the probability of which is low. The RUT is not recommended for confirming infection eradication, except when gastrointestinal endoscopy is indicated.
Diagnostic tests for Helicobacter pylori.
| Test | Sensitivity (%) | Specificity (%) | Use |
|---|---|---|---|
| Rapid urease test | 80−95 | 97−99 | Requires gastric biopsy. Rapid. Can produce false negatives in the context of use with PPIs, antibiotics, bismuth, or gastrointestinal bleeding.Not recommended for eradication. |
| PCR | 97−100 | 98 | Enables the identification of specific genes of the bacterium and the evaluation of antibiotic susceptibility. Considered by some to be the gold standard. |
| Labeled urea breath test | 96−97 | 93−96 | Very good sensitivity and specificity. PPIs should be suspended 2 weeks prior to the test because they reduce sensitivity.It can be used before and after treatment. |
| Serologic test | 55−100 | 58−96.8 | Varies according to the kit utilized.It can only be used for monitoring eradication. |
| H. pylori stool antigen test | 83 | 87−94 | There are tests that use immune enzyme assays and immunochromatographic assays.Easy to implement.Can be used before and after treatment. |
This breath test utilizes the ingestion of 13C-labeled or 14C-labeled urea. If H. pylori is present, the bacterium’s urease enzyme releases the isotope-labeled CO2, which is measured and compared with a baseline value. In general, sensitivity and specificity are above 90%54. PPIs should be suspended before the test because they reduce its sensitivity. The advantage of the exhaled breath test is that it is noninvasive and can also be utilized to evaluate H. pylori eradication, with a high diagnostic yield55.
SerologyH. pylori serology reveals exposure to the microorganism, with varying sensitivity and specificity values, depending on the serologic kit employed. A limitation of serologic analyses is the fact that they do not detect active infection, and so cannot be used for monitoring therapy56. Serology testing is more useful in population studies on the prevalence of H. pylori infection.
Helicobacter pylori stool antigen testStool antigen tests utilize either the technique of enzyme immunoassay or that of rapid immunochromatography. Some of the tests use monoclonal antibodies and others use polyclonal antibodies that are specific for H. pylori antigens57. The advantage of the stool antigen test is that it is easy to implement at different centers, as well as the fact that the stool antigen has been evaluated in H. pylori eradication control, with good diagnostic yield. Care should be taken in patients with diarrhea because watery stools can decrease the sensitivity of the test.
Molecular testsMolecular tests can be useful in diagnosing H. pylori infection. The most widely used is the polymerase chain reaction, or PCR, which along with detecting bacteria, enables evaluating pathogenic and specific genes for antimicrobial resistance. Conserved genes in the H. pylori bacterium, such as ureA, ureC, 16SrRNA, 23SrRNA, and Hsp60, are utilized to perform PCR58. Another advantage of the technique is that the sample can be extracted from the same biopsy specimen utilized for the RUT. Some studies have shown up to 100% sensitivity and 98% specificity for PCR as a diagnostic method for H. pylori59, by using specific primers for conserved genes in the bacterium.
Antimicrobial resistance and its mechanisms in Helicobacter pyloriUnlike other bacteria that acquire antibiotic resistance through horizontal plasmid transmission, H. pylori has vertical transmission of point mutations involved in the mechanisms of resistance that progressively increases due to the selective pressure exerted by antibiotic use31. The main mechanisms of resistance include mutations in key residues in the proteins the antibiotic binds to, regulation in the transport system or in membrane permeability to reduce antibiotic uptake, increased activity in the oxygen scavengers, and modulation of the function of the enzymes involved in the bacterial metabolism of the drugs52,60,61 (Table 2).
Antibiotic resistance methods of Helicobacter pylori.
| Drug | Mechanism of action61 | Resistance rate in the Americas62 | Mechanism of resistance52 | Mutations detectable through NGS52,60 | Other possible mechanisms52 |
|---|---|---|---|---|---|
| MacrolidesClarithromycin | Bacteriostatic – reversible binding to the V domain of the 23S rRNA peptidyl-transferase, interferes with protein synthesis | 10% |
| rrl codons – 2142, 2142, 2143 |
|
| FluoroquinolonesLevofloxacin | Bactericide –-topoisomerase II inhibition (DNA gyrase and topoisomerase IV) | 15% |
| gyrA codons – 86–88, 91, 97, 130gyrB codons– 463, 481, 484 |
|
| NitroimidazolesMetronidazole | Nitro-anion radicals and imidazole intermediaries that damage subcellular structure and DNA | 23% |
| Not described |
|
| Bismuth | Antimicrobial activity against gastrointestinal pathogens | ? |
| Not described |
|
| TetracyclinesTetracycline | Bacteriostatic – binding to 16S rRNA of the 30S ribosomal subunit, inhibits protein synthesis | 4% |
| 16s rRNA codons – 926−928 |
|
| PenicillinsAmoxicillin | Bactericide – binding to PBPs, interferes with bacterial wall synthesis | 10% |
| PPBP1 codons – 402−403, 555−567 |
|
| RifampicinRifabutin | Bactericide – binding to the DNA-dependent polymerase RNA subunit B, inhibits transcription | 1−2% |
| rpoB codons – 525, 545, 585, 149, 701, 149 |
|
DNA: deoxyribonucleic acid; furR3I: Fur protein; MIC: minimum inhibitory concentration; NGS: next-generation sequencing; OMPs: outer membrane proteins; PBPs: penicillin-binding proteins; recA: bacterial DNA recombination protein; RNA: ribonucleic acid; rRNA: ribosomal ribonucleic acid.
Clarithromycin is a macrolide that binds to the peptidyl-transferase region of the 23S ribosomal ribonucleic acid (23S rRNA) molecule in the 50S ribosomal subunit. Its binding inhibits protein elongation through the premature release of peptidyl-RNA from the acceptor site, blocking protein synthesis52. Resistance to clarithromycin is produced by point mutations in the rrl gene that encodes the 23SrRNA 2142 and 2143 nucleotides63. In descending order of frequency, A2143G, A2142G, and A2142C are the three different mutations found to be present in clarithromycin-resistant H. pylori60. The A2142G and A2142C mutations are associated with high-level cross-resistance to all macrolides, whereas the A2143G mutation is associated with high resistance to erythromycin and intermediate resistance to clindamycin and streptogramine64. There are other less important mechanisms by which H. pylori can acquire resistance to clarithromycin, such as mutations in the HefABC (hp0605/606/607) efflux pump, as well as in the hp1048 (infB) and hp1314 (rpl22) genes52,60,64.
Levofloxacin is a fluoroquinolone whose mechanism of action is the inhibition of DNA gyrase (topoisomerase II) and topoisomerase IV52. Those enzymes are encoded in 4 distinct genes; gyrA and gyrB in the case of DNA gyrase and parC and parE in the case of topoisomerase IV (not present in H. pylori). The mutations in those genes are responsible for the majority of cases of quinolone resistance in bacteria in general65. In the specific case of H. pylori, the mutations in the gyrA gene appear to be the main cause of resistance to fluoroquinolones, and there are also recent reports of mutations in the gyrB gene60. The point mutations of the gyrA gene, in the positions that encode the 86–88, 91, 97, or 130 amino acids, are responsible for fluoroquinolone resistance in H. pylori, and the most frequent are those found at codons 87 and 9152,66,67. In general, there is cross-resistance to the rest of the antibiotics of that family52.
Metronidazole is a prodrug that must penetrate the bacterial cell to be activated. Once inside the cell, flavodoxin or ferredoxin, oxidized with electrons by the pyruvate oxidoreductase complex, reduces the metronidazole nitro group to anionic radicals that inhibit nucleic acid synthesis52,60,63,66. The intracellular environment of H. pylori is microaerophilic; the available oxygen molecule competes with metronidazole for electrons, which favors the formation of superoxides that damage the bacterium’s DNA. Resistance to metronidazole can develop by means of different mechanisms, whether due to an increase in oxygen scavenger activity, a decrease in nitroreductase activity, a decrease in antibiotic uptake, or an increase in enzyme activity52,60. The predominant mechanism of resistance is the decrease in nitroreductase activity and the consequent activation of metronidazole. The mutations that inactivate oxidoreductases, such as rdxA (oxygen-insensitive NADPH nitroreductase), frxA (NADP-flavin oxidoreductase), and fdxB (ferredoxin-like enzyme), directly produce a decrease in the quantity of nitroreductase enzyme present in H. pylori60,63,66. Importantly, metronidazole is one of the few drugs whose resistance can be overcome, given that in vitro resistance does not correlate with in vivo effectiveness, especially when the drug is used as a component of triple or quadruple therapy68.
Amoxicillin is a drug from the beta-lactam group, whose mechanism of action is the binding with penicillin-binding proteins (PBPs) to inhibit bacterial wall synthesis52,60. In the context of its use for treating H. pylori infections, it is important to emphasize that the antimicrobial activity of the drug is pH-dependent and its MIC varies in relation to its ambient pH69. Even though β-lactamase-like genes have been found in the H. pylori genome, no significant β-lactamase activity has been found in the amoxicillin-resistant strains52,60,61. The most important mechanism of resistance to amoxicillin is through point mutations in the genes that encode the PBPs, mainly mutations in the pbp1A gene that lead to the substitutions of amino acids in the PBP1 transpeptidase region61. The absence of the gene that encodes for PBP4 has also been associated with amoxicillin resistance63,64,70. A secondary mechanism, by which some H. pylori strains have an increased MIC to amoxicillin, is through decreased membrane permeability to the antibiotic60. Mutations in the porin protein-encoding hopB and hopC genes cause amoxicillin resistance and have a synergic effect with mutations in PBPs65,71.
The tetracyclines are bacteriostatic antibiotics that bind to the 30S ribosomal subunit, inhibiting protein synthesis52,60. The main mechanism of resistance to this group of antibiotics is through the mutations in the helix 31 region of the 16S rRNA molecule, specifically at positions 926–928, the tetracycline binding site64,72. A second mechanism of resistance described in tetracycline resistant strains is the presence of efflux pumps, which reduce the intracellular concentrations of tetracyclines52,63.
Rifabutin inhibits the transcription of RNA by binding to the RNA polymerase β-subunit that is dependent on RpoB DNA and encoded by the rpoB gene73. Resistance to the drug is due to point mutations of the rpoB gene in four different regions, whether at codon 525–545, 585, 149, or 701. The most commonly found mutations are the first two just mentioned52,61,65,73. The mutations are confined to the areas near the active site of RpoB, where the antibiotic binds, impeding interaction of the enzyme with the drug52–73.
Two mechanisms associated with multidrug-resistant H. pylori are biofilm formation and efflux pumps74,75. The RND family pumps are particularly relevant because they are associated with resistance to different classes of medications. H. pylori has a group of 4 genes considered RND family pumps, which are Hp0605-0607 (hefABC), Hp0969-0971 (hefDEF), Hp1327-1329 (hefGHI), and Hp1487-1489. The YajR homologue TetA (HP1165) has also been found to be involved60,75. Regarding the biofilms formed by H. pylori, they are bacterial communities enclosed in a multidimensional matrix of extracellular polymeric substances that have been tied to chronic infections and a reduced susceptibility to multiple classes of antibiotics28,74. Both mechanisms produce a nonspecific resistance phenotype for a family of drugs and they have been found to be synergistic, given that the H. pylori biofilms have increased efflux pump expression74.
Novel therapeutic options and future perspectivesNew treatments for H. pylori infection are based on meta-analyses of retrospective studies on drugs in selected populations, until finding a regimen with the higher eradication rate, regardless of the percentage. In accordance with the guidelines, the regimens that provide superior percentages are the ones that are considered the treatment status quo, which has impeded the development of new therapies for years. Historically, the development of antibiotics is not profitable for drug companies, given that it implies years of research, the regimens are short (7–14 days), and there is the risk of the rapid development of resistance, invalidating the new drug as an option. And so, novel therapeutics are under investigation, among which are the development of new drugs, vaccines, probiotic use, and nanotechnology.
Of the novel strategies for fighting resistance, an attempt has been made to optimize the factors that influence both the bacterium and the host, to obtain the best results. Antibiotic efficacy lies in the action of the drug in a neutral gastric environment, through gastric acid suppression. In that respect, reference has been made to the use of high doses of second-generation PPIs, given that their liver metabolism is less dependent on CYP2C19 (e.g., rabeprazole or pantoprazole) and they have a longer half-life, albeit the same number of days (3–5 days) are needed to reach their maximum effect in the stomach45. Interestingly, a molecule with a novel mechanism –vonoprazan– was approved in Japan in 2015. It is a selective potassium channel inhibitor that is up to 300-times more selective than traditional PPIs, reaching a therapeutic effect in 1−2 days. It is not affected or influenced by the liver metabolism of the host45. The molecule has been approved by the Japanese Society of Gastroenterology and several meta-analyses propose the combination of amoxicillin and vonoprazan as first-line or second-line therapy for H. pylori infections9. On the other hand, new antibiotics have been developed that are not yet integrated into triple therapy or quadruple therapy regimens for H. pylori. One of them is sitafloxacin, a fourth-generation fluoroquinolone that maintains bactericidal activity at a higher MIC for resistance, even when traditional third-generation quinolones have failed69.
Given that antibiotics are employed to fight the bacterium, the hypothesis that the alteration of the gut microbiota can generate dysbiosis that has a short-term or long-term influence on the therapeutic results and prognosis of the patient emerged3. In the short term, supplementation with probiotics has been attempted. They are bacteria that provide a benefit to the host and can have an influence on H. pylori elimination on their own or through the interaction with the microenvironment51. However, a recent analysis has shown that their supplementation does not improve eradication rates, albeit there could be a beneficial impact by mitigating the adverse effects caused by the combination of the medications76. With respect to the long-term changes in the microbiota, very few bacterial species (viral and fungal) actually manage to survive in the gastric pH. A study in The Lancet, in which a longitudinal one-year follow-up on patients treated with antibiotics was carried out, showed that alterations in the microbiota presented at 14 days and 8 weeks after starting treatment and consistently remitted in the follow-up at one year77, indicating that changes in the microbiota are transitory. However, there is contradictory evidence that a metabolic effect does persist (weight gain, metabolic syndrome), despite the restoration of the intestinal flora, opening the possibility of long-lasting effects after treatment15.
With respect to new technologic developments, the advances in nanotechnology and clinical trials with vaccines should be emphasized. The use of nanotechnology can provide new ways to eradicate bacteria by creating compounds that selectively damage the organism or alter the conditions of its microenvironment. Heavy metal nanoparticles (silver, copper, zinc) binding to a particle that selectively recognizes H. pylori have been shown to be effective in vitro by causing lysis of the microorganism due to alteration of the bacterial wall, but studies that confirm its safety in humans are needed78. Likewise, another strategy is to attempt to prevent H. pylori from forming biofilms. Under conditions of environmental stress, quorum sensing protects the biofilms from the bactericide action of antibiotics and induces the transition of the bacterium to its coccoid form, as well. In that sense, several questions regarding nanotechnology must still be answered74. With respect to vaccines, studies on pediatric populations utilizing H. pylori virulence factors (e.g., ompA, cagA) have failed to demonstrate lasting response and protection against the infection79. Treatment targeted at counteracting virulence factors is still in the early phases but shows promise of heading a new therapeutic revolution51.
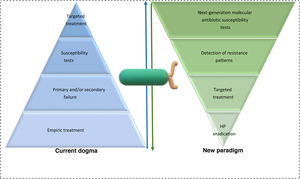
In summary, the development of new therapeutics is important, but we must not lose sight of the horizon. The current problem is not a lack of therapeutic options but rather the inability to establish an epidemiologic surveillance program to know the antibiotic resistance rates, as well as the indiscriminate use of regimens that have no real proven efficacy. At present, the majority of the guidelines prioritize therapeutics that have a higher percentage of effectiveness, and antimicrobial stewardship is relegated to being used when all else fails13,80. More intention-to-treat studies are needed, as well as analyses on the cost-effectiveness and profitability of the next-generation molecular methods, to achieve a change in the international guidelines. The formula needs to be restated; next-generation sequencing (NGS) should be the first step of the algorithm, not the last (Fig. 1).
ConclusionsHelicobacter pylori infection has become a true diagnostic-therapeutic challenge for gastroenterologists. Even though traditional diagnostic methods have acceptable sensitivity and specificity, next-generation molecular diagnostic methods have been shown to be highly sensitive (with close to 100% sensitivity) and specific in the diagnosis of H. pylori. In addition, they have the added advantage of identifying genes that confer antimicrobial resistance, enabling a specific treatment eradication regimen to be prescribed, rather than starting an empiric antimicrobial regimen that the bacterium could be resistant to, as is the current practice.
Furthermore, it is indispensable to consider all the factors associated with infection eradication failure, including those associated with the bacterium, those associated with the host, and importantly, those associated with the healthcare system. It is of the utmost significance for the clinician to clearly and concisely transmit to the patient the treatment to follow, advising him/her ahead of time about possible adverse effects and making digital tools available that serve to maintain adherence to the prescribed treatment.
The increase in antibiotic resistance rates has resulted in a growing decrease in H. pylori eradication rates, producing a direct impact on the quality of life of patients, as well as on the costs of medical care. The diagnostic methods described herein are an innovative strategy for developing an antibiotic stewardship program for eradicating the bacterium. A susceptibility-guided eradication approach, through the detection of mutations in the sensitivity of first-line antibiotics, is the course of the future for achieving a change in the H. pylori treatment paradigm.
Ethical considerationsNo clinical research was conducted to produce the present narrative review, thus neither informed consent nor ethics committee approval were requested.
Financial disclosureNo specific grants were received from public sector agencies, the business sector, or non-profit organizations in relation to this narrative review.
Conflict of interestThe authors declare that there is no conflict of interest.
Author contributionsGarrido-Treviño, López-Martínez, Tijerina-Rodríguez, and Flores-Hinojosa drafted the article and equally contributed to the formulation of the tables and images. Bosques-Padilla reviewed the draft and wrote the final document.
Please cite this article as: Garrido-Treviño LF, López-Martínez M, Flores-Hinojosa JA, Tijerina-Rodríguez L, Bosques-Padilla F. Tratamiento empírico vs tratamiento basado en susceptibilidad para erradicar H. pylori: ¿es posible cambiar este paradigma usando métodos moleculares modernos? Rev Gastroenterol Méx. 2022;87:330–341.